A simple loop to add multiple layers
Library leaflet with R is well integrated with pipe writing (%>%). Thus, it is really easy to add multiple layers and options in a leaflet widget.
If you want to add a lot of layers in your widget, it is not necessary to provide a “addPolygons” for each of these layers, you can simply use a loop. You will need to previously standardize the names of your layers to make a clear code. Moreover, this also allows to use different group names for different view options.
The leaflet widget below shows an example of multiple layers viewing, added with a loop. The R-code is available on my github.
Discrepancies between fishing quotas and scientific advices areas
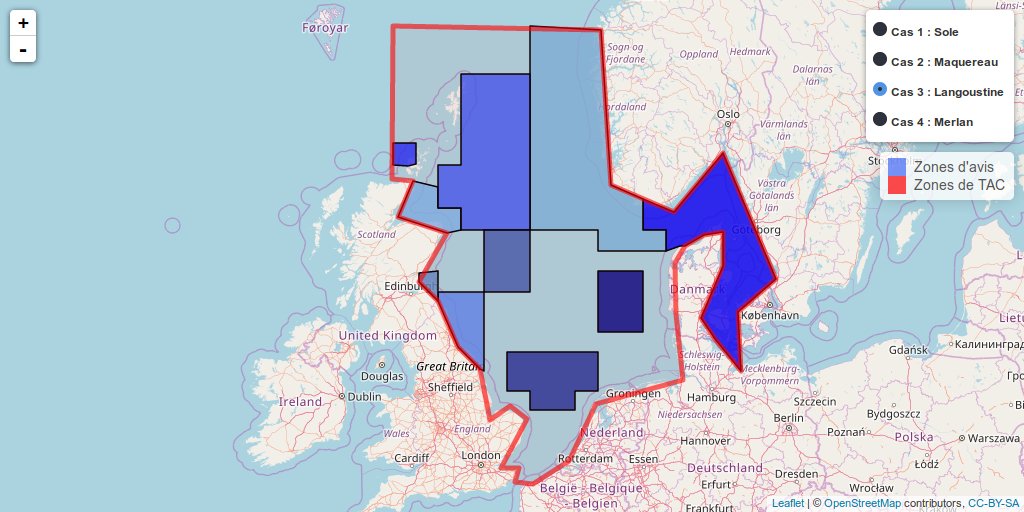
The example below is a widget realised for AFH (French Association of fisheries scientists). This shows different cases for the discrepancies between fishing quotas areas defined by the European commission and areas delimited by scientists to give their advices concerning fish resources. The complete article can be directly downloaded on the AFH website (in French).
Key points of the AFH note
(Translation from French)
- On December, 12-13. 2016, EU ministers of fisheries will set 2017 fishing quotas, in theory according to scientific advice and with the objective of sustainable fisheries that provide maximum sustainable yield.
- Scientific advice is established at the scale of fonctional units for fish populations, which are based on biological knowledge and evolve in time according to new knowledge on species ecology.
- On the other hand, management areas, for which quotas are set, were defined in 1982, and hardly evolved since then.
- Consequently, there is an increasing mismatch between the scale at which advice is given and the scale at which fishing is managed. This results in a very complex and often nontransparent quota-setting process for all stakeholders.
- More importantly, these discrepancies create risks for the sustainability of biological resources. They jeopardize the achievement of maximum sustainable yield for the stocks concerned.
Leaflet widget
- Case 1: Concordance management unit - functional unit
- Case 2: Scientific advice is split into several Total Allowable Catches (TACs)
- Case 3: A TAC gathers several advice areas
- Case 4: Complete mismatch between the management unit and the scientific advisory unit (division and / or recombination of advices areas, mixing of previous cases)
R-code to loop multiple layers in a leaflet widget
library(leaflet)
library(rgdal)
# Working directory ----
wd <- ""
# Read multiple shapefiles with standardised name ----
species <- c("Sole", "Maquerel", "Langoustine", "Merlan")
Groupnames <- c("Cas 1 : Sole", "Cas 2 : Maquereau", "Cas 3 : Langoustine",
"Cas 4 : Merlan")
for (sp in species) {
for (type in c("Avis", "TAC")) {
files.sp <- readOGR(dsn = wd, layer = paste(sp, type, sep = "_"),
verbose = FALSE)
assign(paste(sp, type, sep = "_"), files.sp)
}
}
# Create leaflet widget --------------------------------------------------------
m <- leaflet() %>%
addTiles(group = "OSM (default)")
# Add multiple layers with a loop ----------------------------------------------
for (sp.N in 1:length(species)) {
sp <- species[sp.N]
for (type in c("Avis", "TAC")) {
tmp <- get(paste(sp, type, sep = "_"))
# Define different colors depending on type of data
if (type == "Avis") {
myPal <- colorRampPalette(c("blue", "skyblue", "navyblue"))
factpal.Div <- colorFactor(myPal(length(tmp$id)), tmp$id)
m <- m %>%
addPolygons(data = tmp,
fillColor = ~factpal.Div(tmp$id),
color = "#000000",
opacity = 1,
fillOpacity = 0.8,
stroke = TRUE,
weight = 1.5,
smoothFactor = 0.2,
popup = paste0("Zone d'avis : ", tmp$id),
group = Groupnames[sp.N]
)
} else {
myPal <- colorRampPalette(c("red", "orange", "brown"))
factpal.Div <- colorFactor(myPal(length(tmp$id)), tmp$id)
m <- m %>%
addPolygons(data = tmp,
fillColor = "#FF0000",
color = ~factpal.Div(tmp$id),
opacity = 0.6,
fillOpacity = 0.05,
stroke = TRUE,
weight = 5,
smoothFactor = 0.2,
popup = paste0("Zone de TAC : ", tmp$id),
group = Groupnames[sp.N]
)
}
}
} # end of species
# Additional leaflet options ---------------------------------------------------
m <- m %>%
# Add layers controls
addLayersControl(
baseGroups = Groupnames,
options = layersControlOptions(collapsed = FALSE)
) %>%
# Add common legend
addLegend(colors = c("#4367F4", "#FF0000"),
labels = c("Zones d'avis", "Zones de TAC"),
opacity = c(0.8, 0.5))
# Print the map ----------------------------------------------------------------
mCitation:
For attribution, please cite this work as:
Rochette Sébastien. (2016, Dec. 01). "Add multiple layers in leaflet widget with R". Retrieved from https://statnmap.com/2016-12-01-multiple-layers-leaflet-widget-with-rstat/.
BibTex citation:
@misc{Roche2016Addmu,
author = {Rochette Sébastien},
title = {Add multiple layers in leaflet widget with R},
url = {https://statnmap.com/2016-12-01-multiple-layers-leaflet-widget-with-rstat/},
year = {2016}
}